Using driftplots#
driftplots can be used to generate static Matplotlib figures or an interactive viewer (built on Qt).
In the interactive viewer, clicking a spike will display the template (unscaled, whitened) for
the template assigned to that spike.
Below we will cover the main ways to use driftplots.
See the API Reference for a full list of arguments.
Note
See here for a glossary of key terms.
Inputs#
driftplots accepts a path to Kilosort’s output or a SpikeInterface SortingAnalyzer as input.
If passing a path directly to a sorting output, Kilosort 1-4 are supported.
See here for details on how spike amplitudes,
depths and unit templates are computed across Kilosort versions). spike_templates.npy
is used to assign the template for each spike. This follows Kilosort’s original per-spike
template assignment and therefore will not reflect later merges or splits made in Phy.
If passing a SortingAnalyzer, it is expected that the required extensions
have already been computed. See
this example
for the required extensions. Note that the number of spikes displayed will depend
on the argument set for max_spikes_per_unit used when computing "random_spikes".
By default, the number of spikes displayed will be decimated to around 100,000.
Warning
driftplots was designed and tested with Neuropixels probes, but it should also work with most other probe types.
Interactive Viewer#
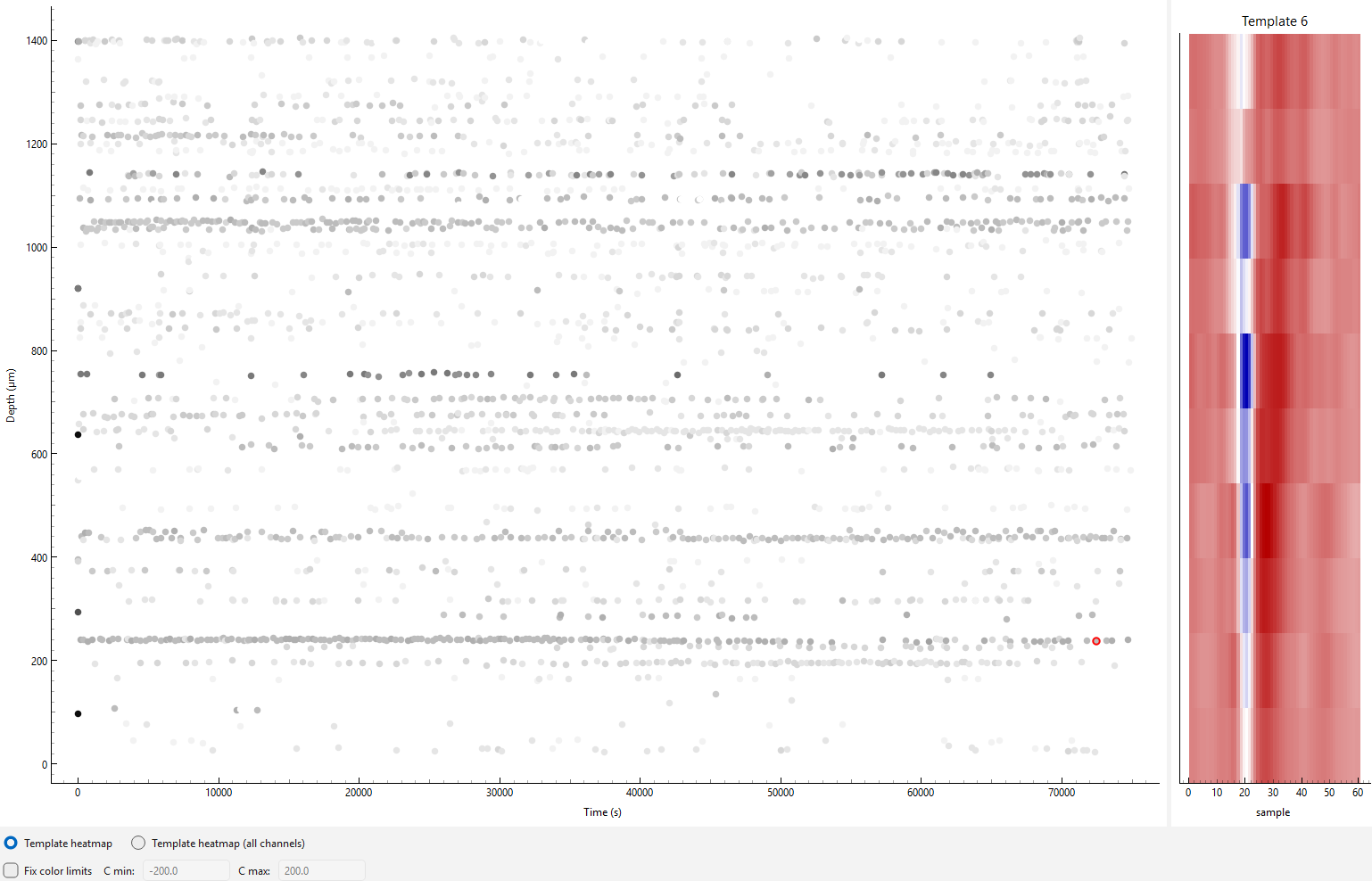
drift_map_plot_matplotlib() generates an interactive viewer
allowing selection of individual spikes on the driftmap. Once selected, the template for that
spike will be displayed on the right-hand side.
from driftplots import DriftPlotter
plotter = DriftPlotter(
"/path/to/sorter_output",
)
driftmap = plotter.drift_map_plot_interactive(
exclude_noise=True,
)
driftmap.plot()
The displayed templates are whitened and are not scaled per-spike i.e. the template will appear the same for all spikes assigned to the same template. The template is whitened and is not scaled to each individual spike. This approach was chosen for two main reasons:This is due to the difficulty of reconstructing individual waveforms from
it is not always possible to reconstruct the waveforms or their amplitudes across kilosort versions (see here)
the main purpose of the interactive mode is to check that templates are identifiably similar across sessions, to ensure little drift has occurred. This is easier with templates rather than noisier spike waveforms.
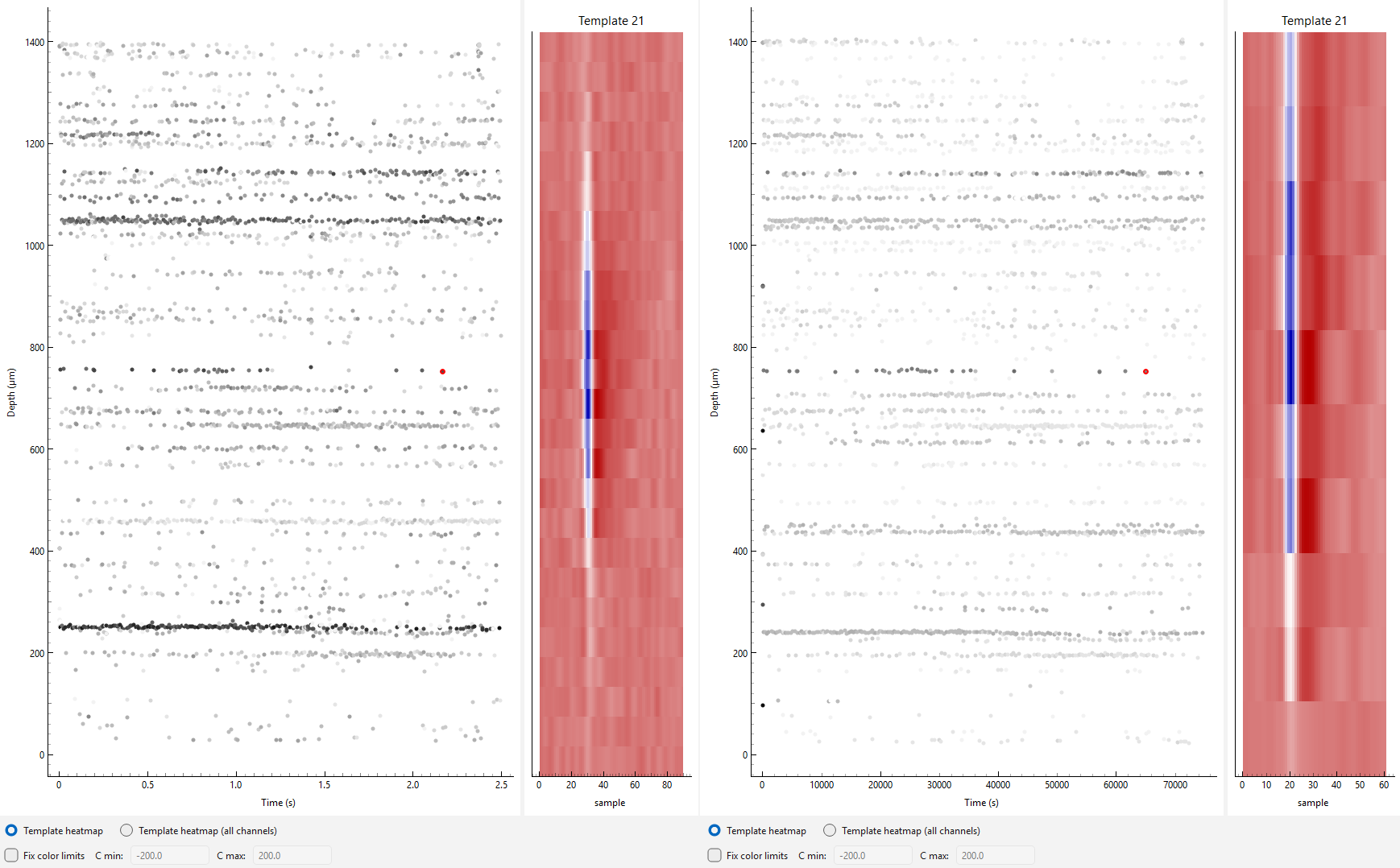
Interactive Viewer with Multiple Plots#
MultiSessionDriftmapWidget can be used to display
multiple interactive plots at once. In this mode, the y-axis
zoom is linked across plots.See the amplitudes section for details
on how to ensure amplitude scaling is consistent across sessions.
import spikeinterface as si
from driftplots import DriftPlotter, MultiSessionDriftmapWidget
# Load the data. In this example we load as a sorting analyzer
# or from the raw kilosort output to demonstrate both methods
data_path = "/path/to/example_data"
analyzer = si.load_sorting_analyzer(data_path / "analyzer.zarr")
sorting_output_path = data_path / "sorting" / "sorter_output"
# Create a list of interactive plots, and collect them
# into a single plot using MultiSessionDriftmapWidget
panels = []
for title, path_or_analyzer in zip(
["Session 1", "Session 2"],
[analyzer, sorting_output_path]
):
plotter = DriftPlotter(path_or_analyzer)
plot = plotter.drift_map_plot_interactive()
panels.append(plot)
multi = MultiSessionDriftmapWidget(panels)
multi.plot()
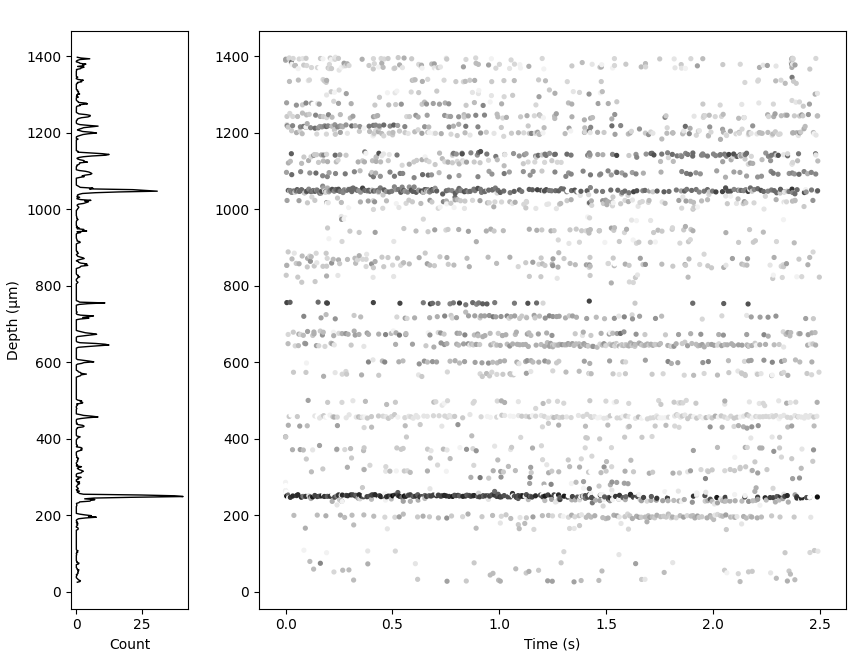
Matplotlib Mode#
drift_map_plot_matplotlib() returns a static Matplotlib figure. It
takes all the same arguments as the interactive viewer
but can additionally plot a 1D activity histogram next to the driftmap.
See this example for how to stitch Matplotlib figures together across an experimental project to quickly assess recording quality and stability.
import matplotlib.pyplot as plt
import spikeinterface as si
from driftplots import DriftPlotter
analyzer = si.load_sorting_analyzer("/path/to/analyzer.zarr")
plotter = DriftPlotter(analyzer)
fig = plotter.drift_map_plot_matplotlib(
exclude_noise="KSLabel",
add_histogram_plot=True,
weight_histogram_by_amplitude=True,
)
plt.show()
Aligning Amplitudes Across Sessions#
driftplots provides options for excluding spikes based on their
amplitudes and scaling the color of the scatter points by amplitude.
This can be useful when a small number of high- or low-amplitude spikes dominate
the color scaling, with a few very light/dark spots with the rest grey.
As such it is useful to apply the same amplitude filtering and colormap
scaling to all plots when comparing multiple sessions. get_amplitudes()
can be used to pool amplitudes across sessions, allowing cutoffs to be calculated
across all sessions and applied to all plots.
import numpy as np
import spikeinterface as si
from PySide6 import QtWidgets
from driftplots import DriftPlotter, MultiSessionDriftmapWidget, get_amplitudes
analyzer = si.load_sorting_analyzer("/path/to/an/analyzer.zarr")
SORTING_SESSIONS = [
"/path/to/a/sorting",
analyzer
]
all_spike_amplitudes = get_amplitudes(
SORTING_SESSIONS, exclude_noise=False, concatenate=True
)
min_cutoff, max_cutoff = np.percentile(all_spike_amplitudes, (0, 95))
app = QtWidgets.QApplication([])
panels = []
for path_or_analyzer in SORTING_SESSIONS:
plotter = DriftPlotter(path_or_analyzer)
plot = plotter.drift_map_plot_interactive(
filter_amplitude_mode="absolute",
filter_amplitude_values=(min_cutoff, max_cutoff),
amplitude_cmap_scaling=(min_cutoff, max_cutoff),
n_color_bins=25,
)
panels.append(plot)
multi = MultiSessionDriftmapWidget(panels)
app.exec()